Impact of Whole Genome Sequencing in Healthcare
Genome sequencing services are increasingly being utilized by physicians and their patients in both research and clinical settings. Developing standards and procedures for the use of sequencing information in clinical medicine is an urgent need with numerous obstacles to integrity and storage of sequencing data, interpretation, and responsible clinical integration.
 The MedSeq Project, funded by the NIH, was the very first study exploring the use of whole genome sequencing (GS) in both a healthy population and a population with suspected genetic cardiac disease. The MedSeq Project was designed to explore the medical, behavioral and economics impacts of incorporating GS into everyday medicine. We examined how providers with and without genetics training communicated genetic information to their patients and how patients and providers responded to this genetic information. We also explored the downstream impact this testing had on the health system at large.
The MedSeq Project, funded by the NIH, was the very first study exploring the use of whole genome sequencing (GS) in both a healthy population and a population with suspected genetic cardiac disease. The MedSeq Project was designed to explore the medical, behavioral and economics impacts of incorporating GS into everyday medicine. We examined how providers with and without genetics training communicated genetic information to their patients and how patients and providers responded to this genetic information. We also explored the downstream impact this testing had on the health system at large.
Our research team of more than 50 investigators, staff and expert consultants spent over a year designing a protocol to explore the integration of WGS in clinical medicine, which obtained Institutional Review Board approval in August 2012. We enrolled 11 primary care physicians (PCPs) and 102 of their generally healthy middle-aged patients ages 40-65 to evaluate the use of General Genomic Medicine, and 9 cardiologists and 100 of their patients with hypertrophic or dilated cardiomyopathy to evaluate the use of Disease-Specific Genomic Medicine.
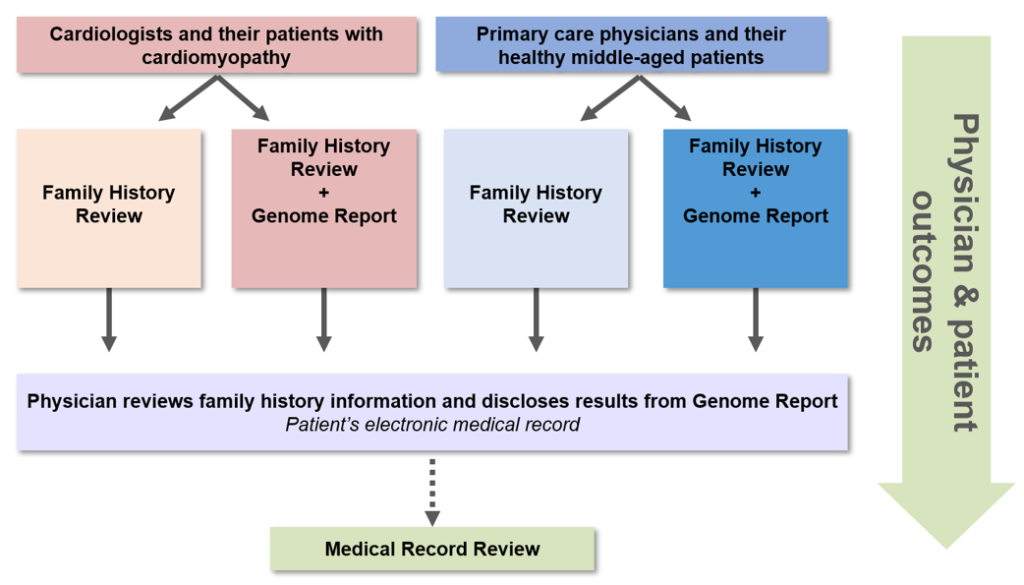
We randomized physicians and their patients within each of the above models to receive clinically meaningful information derived from WGS plus a family history assessment versus current standard of care plus a family history assessment without the use of WGS. The team designed an educational module for the enrolled physicians to complete prior to disclosing WGS results to their patients, including two in-person lectures and four hours of online coursework (accredited by the Partners HealthCare System for a maximum of 6.5 AMA PRA Category 1 Continuing Medical Education Credit(s)). In addition, we created a MedSeq Project Genome Resource Center (GRC), staffed with medical geneticists and genetic counselors, for the physicians to consult regarding their patients’ Genome Report and/or family history information.

Our team designed a mixed-methods approach, utilizing quantitative surveys and qualitative methods, to examine preferences and motivations of physicians and patients enrolled, evaluate the flow and utilization of genomic information within the clinical interactions, and assess understanding, behavior, medical consequences and healthcare costs associated with the use of WGS in these models of medical practice.
For more detailed information about the MedSeq Project, including study design and outcome measures, visit our clinical trial page:
For scientific papers and media coverage, or to donate to our MedSeq Project, visit the links below or email us at g2p@bwh.harvard.edu.